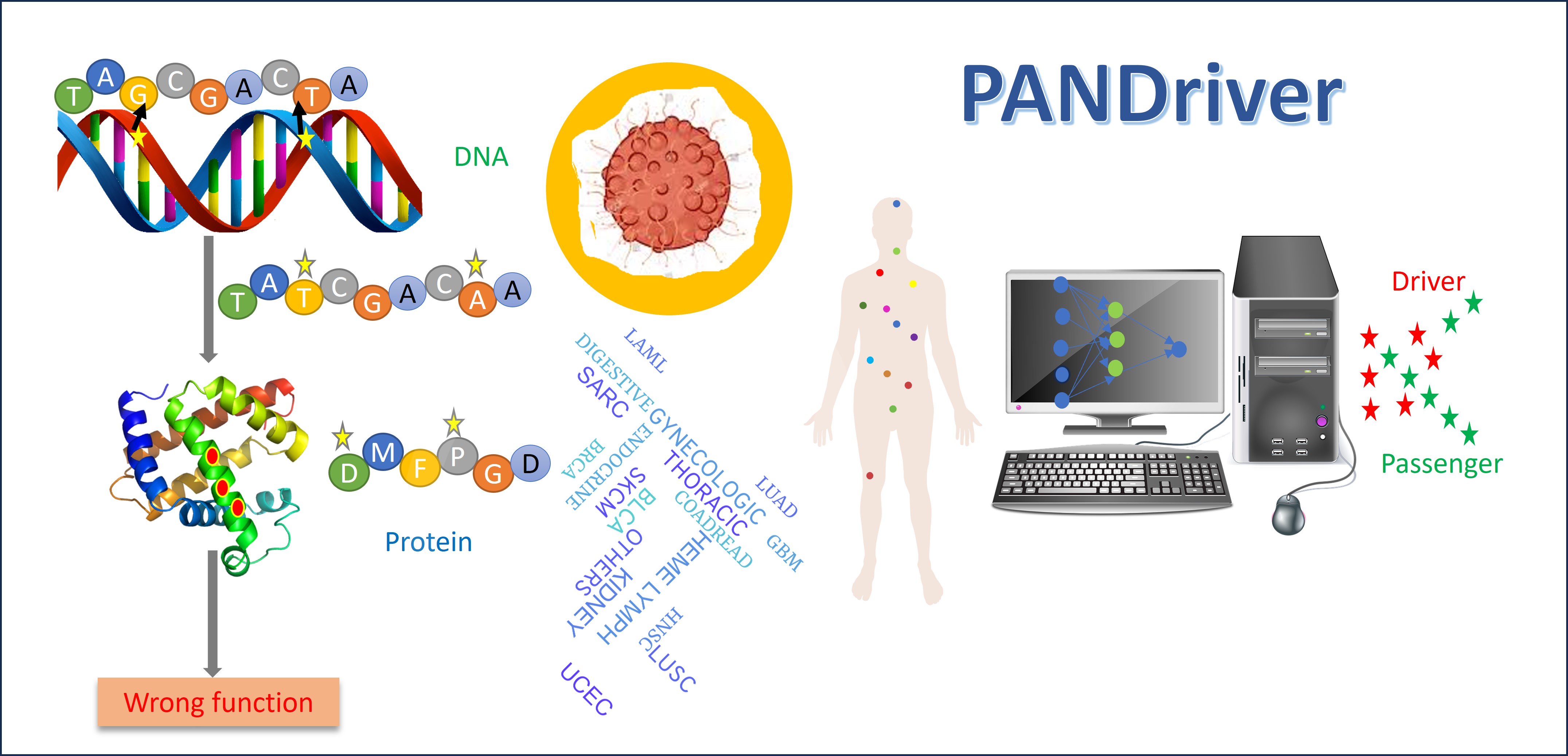
We have developed a method for identifying driver mutations across 15 different cancer types. To develop this method we used amino acid sequence based features (Ex: physicochemical properties, conservation scores, predicted secondary structure and peptide motifs), structure-based properties (Ex: residue depth, accessible surface area (ASA)), and network based properties (Ex: centralities) to discriminate the driver mutations from the passengers. This method used AlphaFold structures for deriving structure based properties. This method will help in prioritizing driver mutations from the pool of mutations obtained from experiments.